Our Technology
Role for hyaluronan in making primal mesenchymal stromal cells - Stromacytes
Our technology is based on our research on the role of hyaluronan in stem cell differentiation. This is a sugar polymer with wide spread roles in organ development, function and disease that depend on how it is presented to cells and other factors that interact with it. In our most recently peer reviewed study published in Stem Cell Research & Therapy, we report how it is sufficient to make distinctive primal mesenchymal stromal cells, we call Stromacytes, capable of modifying the function of blood cell lineages in the same way as adult organ derived cells.
Further investigations on Stromacytes
To understand how hyaluronan exerts its effect on stem cell differentiation we have deeply characterised cell gene expression following its presentation by our original and new methods versus the state of the art. This has informed distinctive changes induced by hyaluronan and co-factors, the nature of the primal cells which result, and the identity of molecules and pathways involved with potential to mediate regeneration of cells and tissues in ageing and adult onset diseases (Figure).
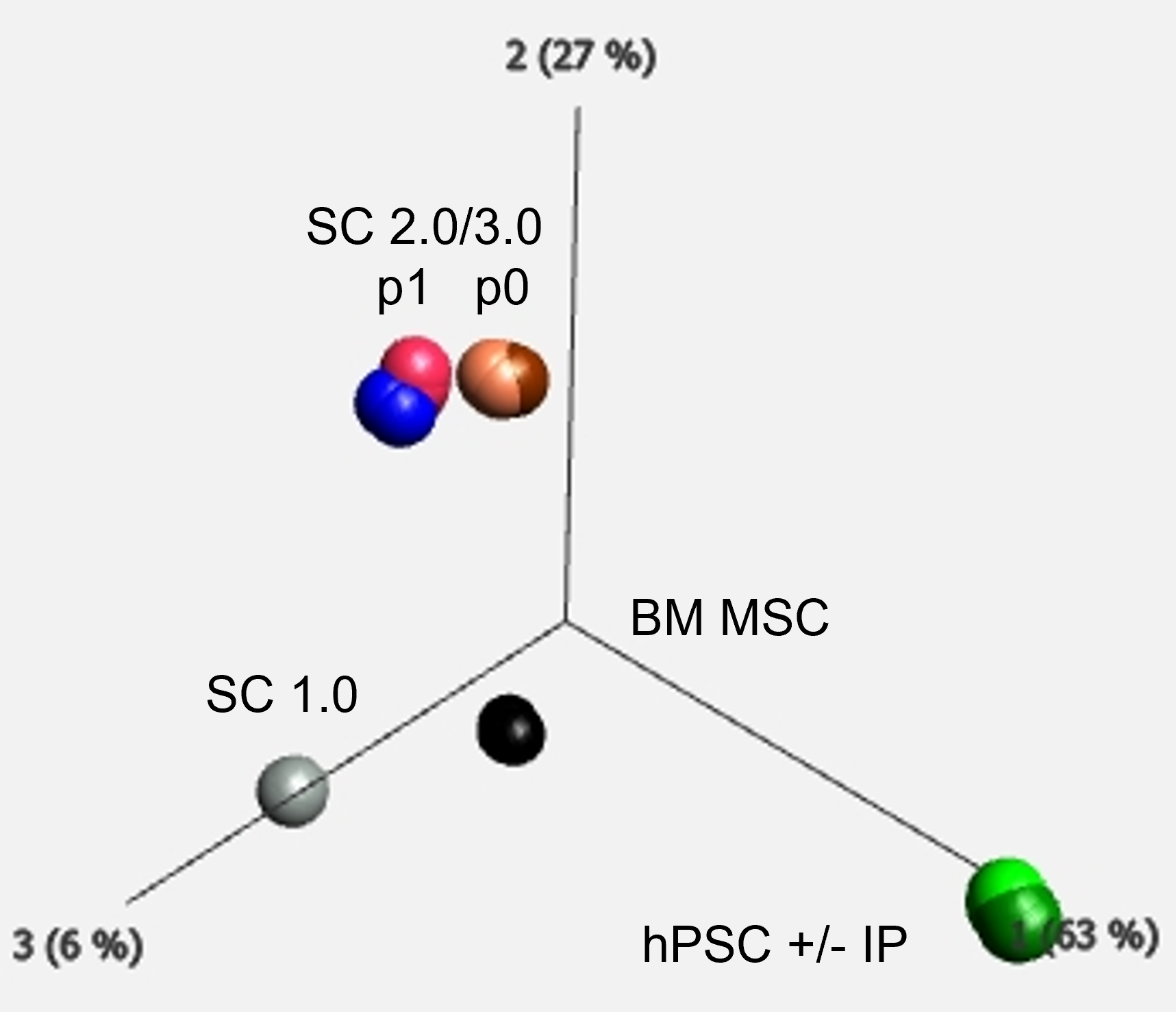
Characterisation of cell gene expression by Principal Component Analysis of all expressed genes assessed by Total RNA sequencing in Stromacyte variants (SC 1.0, 2.0, 3.0) at designated passage (p) in culture, differing in how hyaluronan is presented by our original stromacytes (1.0) and applying new intellectual proprietary (IP) to make alternative stromacyte variants (SC3.0) applied to human pluripotent stem cells (hPSC + IP) vs a comparable state of the art method (SC2.0 and hPSC - IP). These were also compared to human bone marrow mesenchymal stromal cells (BM MSC). Analysis performed using Qlucore Informatics Tools.
Stromacyte release proteins active in healing
In further investigations we have used Liquid Chromatography Mass Spectrometry (LCMS) and proteomic analysis to compare proteins released by variations of primal cells (Stromacytes) and adult bone marrow mesenchymal stromal cells into culture medium. This revealed that although there were differences in relative enrichment between cells, the same 400+ proteins appeared to be secreted by all cells. Bioinformatics analysis associated the activity of these proteins with molecular pathways involved in modulating immune cell function and extracellular matrix organisation involved in tissue healing (Figure).
Gene ontology network of Biological Process Pathways common to 400+ proteins released by stromocyte variants (SC1.0, 2.0, 3.0) and BM MSC in minimal chemically defined medium conditioned for 24 hrs . Each protein is represented by two or more unique peptides detected by LCMS. Two pathways (nodes) are connected if they share 20% or more genes (proteins). Solid nodes are more statistically significantly enriched protein sets. Bigger nodes represent larger protein sets. Thicker edges represent more overlapped proteins. Gene Ontology networks calculated using ShinyGO 0.76
Released proteins enriched in Stromacytes
Variant primal stromacytes made by our hyaluronan based induction technology differ from each other and adult bone marrow mesenchymal stroma cells by relative enrichment of released proteins. One variant is enriched for biological activity in cell, tissue and immune system formation and functions important in healing with ageing. So, primal cells have potential to mimic adult tissue mesenchymal stromal cells, and more functions supporting cell and tissue regeneration (Figure).
Proteomic analysis of proteins released by cells in culture. (A) Principal component Analysis of 400+ proteins secreted by three independent samples of each Stromacyte variant (SC 1.0, 2.0, 3.0) vs bone marrow (BM) mesenchymal stromal cells. This shows distinctiveness of released protein profile for each type of cell as a whole. (B) Heat map of hierarchically clustered proteins showing enrichment for specific proteins (rows) in three independent biological replicates of each cell type (columns). Map is normalised to a mean equal to zero and variance equal to 1, Intensity of squares for each protein in each sample reflect log2fold increase (yellow) or decrease (blue) relative to mean (black). (C) Biological process pathways enriched in SC 3.0 released proteins. Two pathways (nodes) are connected if they share 20% or more proteins. Solid nodes are more significantly enriched protein sets. Bigger nodes represent larger proteins sets. Thicker edges represent more overlapped proteins. PCA and Heat Map prepared using Qlucore Informatics Tools. Gene Ontology networks calculated using ShinyGO 0.76
Applications & Development
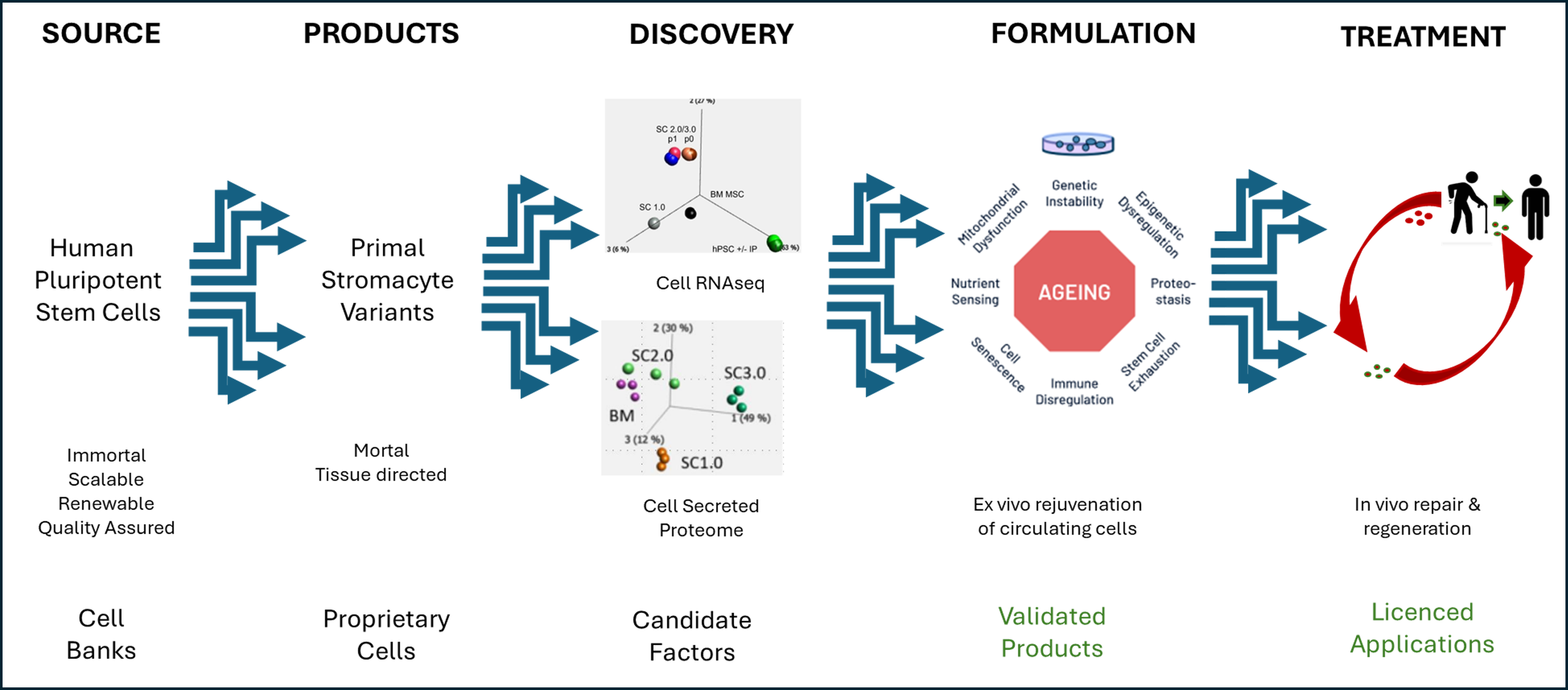
Our goal is to promote healthy ageing and sustainable care of chronic diseases, by applying our expertise and assets in the preclinical discovery, validation and licensed supply of candidate cell rejuvenating & health regenerative formulations of cells and factors.
The road to innovating new therapeutic interventions is long and complex requiring multi-disciplinary expertise and commitment. We welcome opportunites to contribute to new and ongoing initiatives with the capabilities we can offer.